A01-1 Multi-scale theory of genome modality
As only one theoretical research project in the planned research groups of “Genome Modality”, we collaborate with experimental groups explaining measured data via physical models, through which we deepen biological knowledge on genome modality. As a physical basis, we elucidate hierarchic structures of DNA, nucleosomes, chromatin, and chromosome, via multiscale physical theories and simulations. Furthermore, we study mechanisms of chromosome structure formation at interphase and mitosis, by physical modeling and simulations of SMC proteins (cohesin and condesin). Specifically, we have the following four goals:
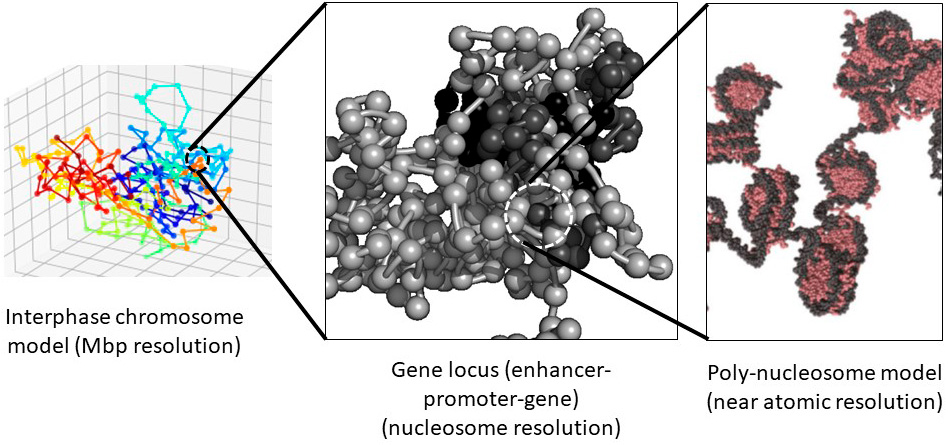
1) To elucidate hierarchic structures of DNA, nucleosomes, chromatin, and chromosome.
2) To build physical models of chromosome formation via SMC proteins.
3) To construct theoretical models for the coupling of chromatin structure and transcription.
4) To uncover physics of non-nucleosomal chromatin condensations.
We also contribute to make an integrated tool, “Genome Modality Suite”, which collects outcomes from experimental groups and facilitates collaborations within the research area.
-

Shoji Takada Project Leader
- Graduate School of Science, Kyoto University
- http://theory.biophys.kyoto-u.ac.jp/index_e.php
-

Yukitaka Ishimoto Co-Project Leader
- Faculty of Systems Science and Technology, Akita Prefectural University
- http://www.akita-pu.ac.jp/system/mise/thermal_fluid/fluid/en/
-

Takahiro Kenmotsu Co-Project Leader
- Faculty of Life and Medical Sciences, Doshisha University
- https://dmpl.doshisha.ac.jp/?lang=en
-

Tetsuya Yamamoto Co-Project Leader
- Institute for Chemical Reaction Design and Discovery
- https://www.icredd.hokudai.ac.jp/yamamoto-tetsuya/

